Non-coding RNAs & RNA interference
 RNA molecules that do not encode protein account for a surprisingly large proportion of transcription in complex organisms including humans. We now know that at least some of these non-coding (nc)RNAs have regulatory functions, representing a previously hidden layer of complexity in genome regulation. Two broad categories of regulatory ncRNAs can be distinguished: long ncRNAs, which function without significant processing, and short ncRNAs that are processed from longer precursors by the RNA interference (RNAi) pathway.
RNA molecules that do not encode protein account for a surprisingly large proportion of transcription in complex organisms including humans. We now know that at least some of these non-coding (nc)RNAs have regulatory functions, representing a previously hidden layer of complexity in genome regulation. Two broad categories of regulatory ncRNAs can be distinguished: long ncRNAs, which function without significant processing, and short ncRNAs that are processed from longer precursors by the RNA interference (RNAi) pathway.
RNAi is a gene silencing mechanism conserved throughout eukaryotes. It is characterised by small ncRNA intermediates that provide the specificity to direct silencing effector complexes to homologous target sequences. Divergent RNAi pathways involve different types of small RNA and direct silencing in different ways; for example, micro (mi)RNAs typically repress expression at the level of translation, whereas short interfering (si)RNAs can mediate either transcript cleavage, or transcriptional repression via chromatin modification.
Chromatin & Epigenetics
 In eukaryotic nuclei, DNA associates with histone proteins to form chromatin. Remodelling of chromatin, for example through methylation of DNA or of histone tails, can mediate stable changes in gene expression without any change in the DNA sequence, a phenomenon termed epigenetics. Broadly speaking, two distinct chromatin states can be distinguished: transcriptionally active euchromatin, rich in histone acetylation, and compact, transcriptionally repressed heterochromatin, characterised by histone methylation.
In eukaryotic nuclei, DNA associates with histone proteins to form chromatin. Remodelling of chromatin, for example through methylation of DNA or of histone tails, can mediate stable changes in gene expression without any change in the DNA sequence, a phenomenon termed epigenetics. Broadly speaking, two distinct chromatin states can be distinguished: transcriptionally active euchromatin, rich in histone acetylation, and compact, transcriptionally repressed heterochromatin, characterised by histone methylation.
Heterochromatin has important regulatory and structural roles in maintaining genome stability. However, it must be accurately positioned to avoid silencing essential genes. Both long and short ncRNAs have been implicated in specifying target domains for heterochromatin assembly. For example, long ncRNAs such as Xist and AIR mediate chromatin modifications associated with X-inactivation and imprinting in mammals, while RNAi is involved in heterochromatin assembly in a range of organisms including plants, vertebrates, and fission yeast. Defects in such RNA-directed chromatin modification have been linked to a range of human diseases including cancers; however, the molecular mechanisms underlying this process remain poorly understood.
RNA-directed chromatin modification in fission yeast
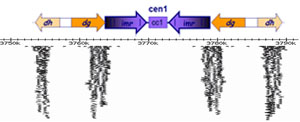
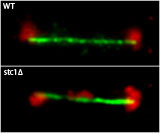 Our aim is to dissect the mechanism and regulation of RNAi-directed chromatin modification. We use the fission yeast Schizosaccharomyces pombe as a model system; this single-celled eukaryote is genetically tractable, yet shares both RNAi and chromatin components with humans. As in higher eukaryotes, the centromeres of fission yeast are flanked by repetitive sequences that are assembled in heterochromatin. Deletion of core RNAi components, including Dicer and Argonaute (present in single copy in S. pombe), results in defects in centromere function due to disruption of this heterochromatin.
Our aim is to dissect the mechanism and regulation of RNAi-directed chromatin modification. We use the fission yeast Schizosaccharomyces pombe as a model system; this single-celled eukaryote is genetically tractable, yet shares both RNAi and chromatin components with humans. As in higher eukaryotes, the centromeres of fission yeast are flanked by repetitive sequences that are assembled in heterochromatin. Deletion of core RNAi components, including Dicer and Argonaute (present in single copy in S. pombe), results in defects in centromere function due to disruption of this heterochromatin.
We currently have a good framework for understanding RNAi-dependent heterochromatin assembly in fission yeast. We know that the pericentromeric repeat sequences of S. pombe are transcribed, producing ncRNAs that are processed by Dicer into short interfering (si)RNAs. These siRNAs are bound by the RNAi effector protein Ago1, resulting in targeting of Ago1 and associated RNAi machinery to homologous nascent transcripts. This RNAi activity is required for recruitment of the Clr4 histone methyltransferase complex (CLRC) to cognate chromatin. Clr4 can then mediate methylation of histone H3 on lysine 9 (H3K9me), promoting binding of the HP1 protein Swi6, and hence heterochromatin formation.
Current Research
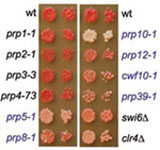
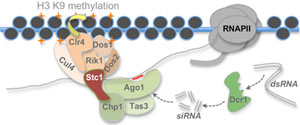 Our current research focuses on understanding how RNAi is integrated with other cellular pathways, including chromatin modification, and how the process is regulated. We previously identified Stc1, a LIM domain protein that associates with both RNAi and chromatin modification components, and appears to play a key role in mediating the RNAi-dependent recruitment of chromatin modifiers to chromatin. We are now dissecting the mechanism by which Stc1 couples RNAi to chromatin modification.
Our current research focuses on understanding how RNAi is integrated with other cellular pathways, including chromatin modification, and how the process is regulated. We previously identified Stc1, a LIM domain protein that associates with both RNAi and chromatin modification components, and appears to play a key role in mediating the RNAi-dependent recruitment of chromatin modifiers to chromatin. We are now dissecting the mechanism by which Stc1 couples RNAi to chromatin modification.
In addition to Stc1, we are investigating the roles of several other proteins that were identified through genetic screens as contributing to heterochromatin assembly in fission yeast. We are also carrying out new genetic screens designed to identify proteins important for proper regulation of RNAi-directed chromatin modification. In the long term, we hope these studies in fission yeast will help us understand a range of human diseases linked to defects in the mechanism or regulation of RNA-directed chromatin modification.


